Machine learning generates pictures of proteins in 5D
Newly developed method does it all with just a few pixels
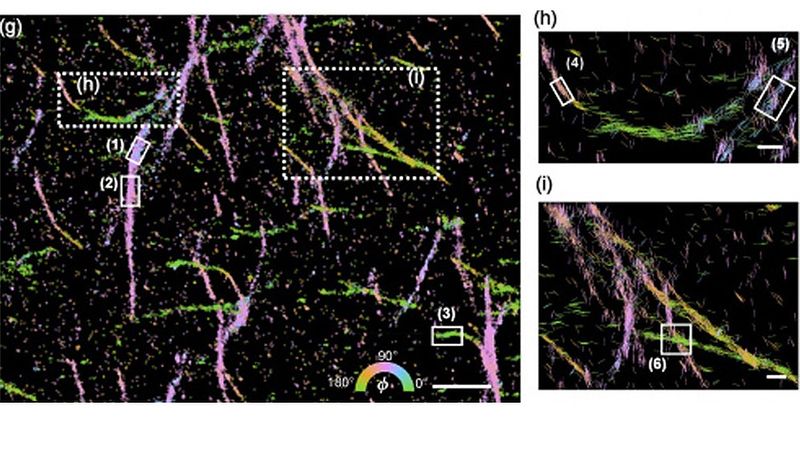
Amyloid proteins. Their lengths and directions indicate the magnitude of their in-plane orientations and their orientations, respectively. (Credit: Lew Lab)
By combining machine learning with the laws of physics, researchers in the lab of Matthew Lew, associate professor in the Preston M. Green Department of Electrical & Systems Engineering at Washington University in St. Louis, have been able to sort out the orientation and position of overlapping single molecules in 5D from a single image.
Their research was published Sept. 26, 2022, in the journal Optics Express.
The five dimensions in question aren’t new, hidden spatial dimensions; instead, a team headed by Tingting Wu, a PhD student in the McKelvey School of Engineering imaging sciences program, was able to design a system that could tell her the orientation of a molecule in 3D space as well as its position in 2D, five parameters from a single, noisy, pixelated image.
For context, a basic computer vision algorithm, for instance, may look at an image and find a person among trees in a forest, Lew said, a task termed classification. “But many algorithms can’t tell you what direction the person is moving, how far away they are, or if that person happens to be inhaling or exhaling—examples of estimation tasks.”
To wrest this additional complex information from a seemingly simple spot of light, the team did design a machine learning algorithm, but added an extra step.
“A lot of people use AI end-to-end,” Wu said. “Just put in the thing you have and ask the neural network to give you the thing you want.” She decided to break the problem into two steps to lighten the load on the algorithm, making it more robust.
The kind of imaging carried out in the Lew lab — of single molecules — tends to be very “noisy,” containing “specks” or fluctuations that can obscure an image. For most machine learning neural networks, Lew said, “robustly dealing with that kind of noise can be very complicated to learn.”
Humans, however, have already learned how signals from the molecules of interest and this noise are combined together within microscope images. Instead of asking the algorithm to re-learn the laws of physics, the team added a second, “post-processing” algorithm–a straightforward computation that applied these physical laws to the results from the first algorithm.
“It’s like I’ve separated two problems into two algorithms,” Wu said.
Using a machine learning algorithm coupled with post-processing allows the Lew lab to uncover the structureon the right from the noisy, pixelated image on the left. The image on the right is color coded with estimated 3D orientation. (Credit: Lew lab)
After processing thousands of snapshots, the result, Wu said, is a “beautiful image” that uses color, curvature and direction to indicate how thousands of molecules are connected to each other.
Ultimately, this system will be able to help researchers better understand biological processes at tiny scales — like the way in which amyloid proteins assemble to form the tangled structures associated with Alzheimer’s disease.
This research was supported by the National Institute of General Medical Sciences (R35GM124858) and the National Science Foundation (ECCS-1653777)
